Search Result
Results for "
RNA structure
" in MedChemExpress (MCE) Product Catalog:
1
Biochemical Assay Reagents
1
Isotope-Labeled Compounds
| Cat. No. |
Product Name |
Target |
Research Areas |
Chemical Structure |
-
- HY-W200545
-
-
-
- HY-103006
-
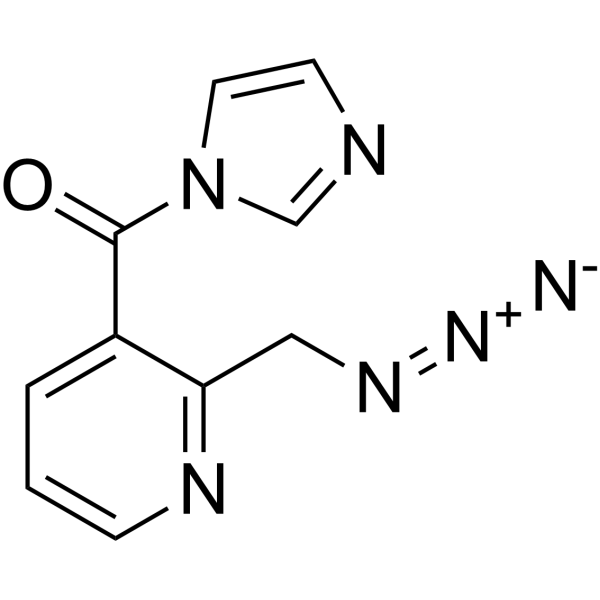
NAI-N3
5 Publications Verification
|
Biochemical Assay Reagents
|
Others
|
|
NAI-N3 is a RNA acylation reagent that enables RNA purification. NAI-N3 is a dual-function SHAPE (selective 2-hydroxyl acylation and profiling experiment) probe (RNA structure probe and enrichment) .
|
-
-
- HY-18408
-
|
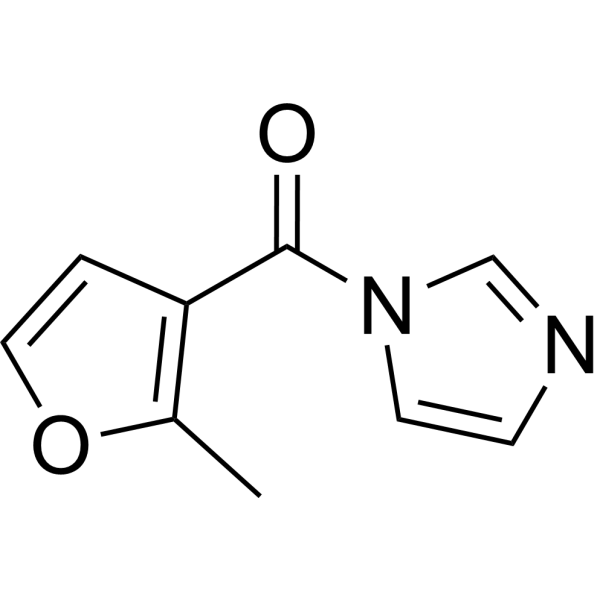
FAI
|
Biochemical Assay Reagents
|
Others
|
|
5S rRNA modificator is a suitable electrophile for 2’-hydroxyl acylation on structured RNA molecules, yielding accurate structural information comparable to that obtained with existing probes; 5S rRNA RNA modification.
|
-
-
- HY-D1408
-
|
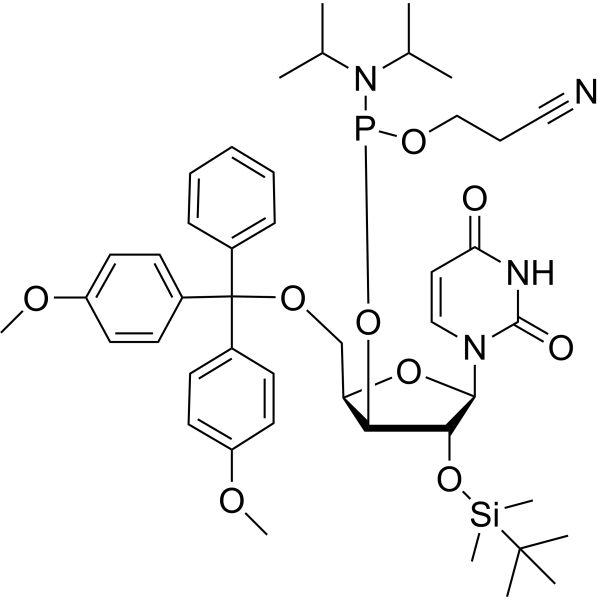
DMTr-4'-Methyluridine-CED-TBDMS phosphoramidite
|
DNA Stain
|
Cardiovascular Disease
|
|
DMTr-4'-Me-U-CED-TBDMS phosphoramidite (DMTr-4'-Methyluridine-CED-TBDMS phosphoramidite), a dye reagent for oligonucleotide labeling, can be used for the research of applications in RNA therapeutics, RNA aptamers, and ribozymes for elucidating RNA structure. DMTr-4'-Me-U-CED-TBDMS phosphoramidite represents a probe with wide utility for elucidation of RNA structure .
|
-
-
- HY-D1409
-
|
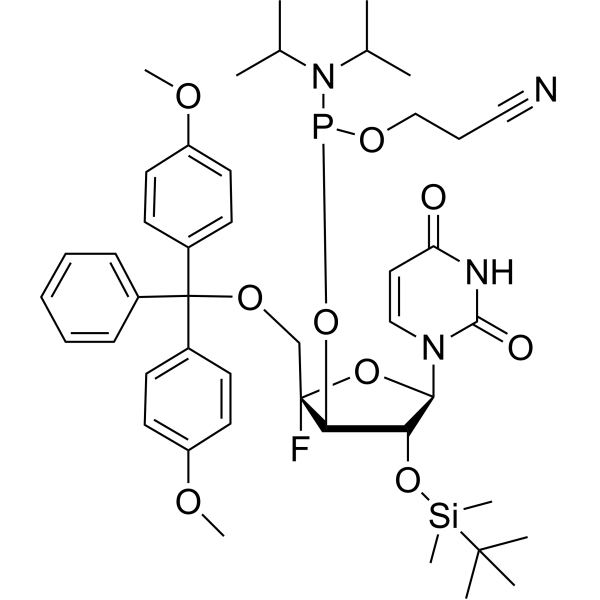
DMTr-4'-F-uridine-CED-TBDMS phosphoramidite
|
DNA Stain
|
Cardiovascular Disease
|
|
DMTr-4'-F-U-CED-TBDMS phosphoramidite (DMTr-4'-F-uridine-CED-TBDMS phosphoramidite), a dye reagent for oligonucleotide labeling, can be used for the research of applications in RNA therapeutics, RNA aptamers, and ribozymes for elucidating RNA structure. DMTr-4'-F-U-CED-TBDMS phosphoramidite represents a probe with wide utility for elucidation of RNA structure .
|
-
-
- HY-18407
-
|
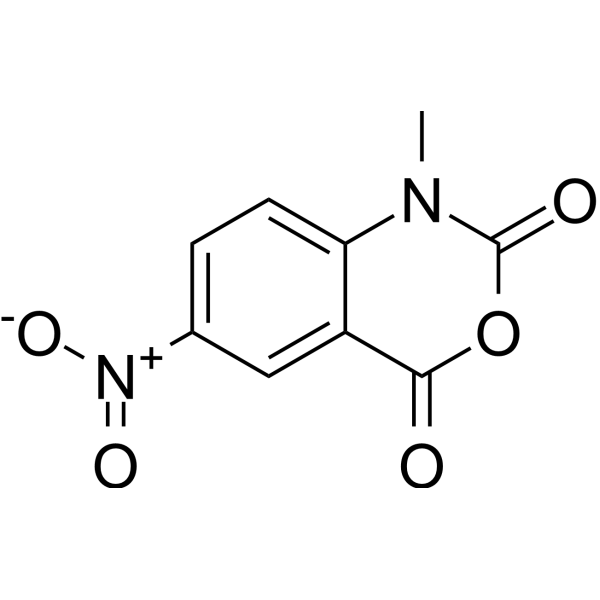
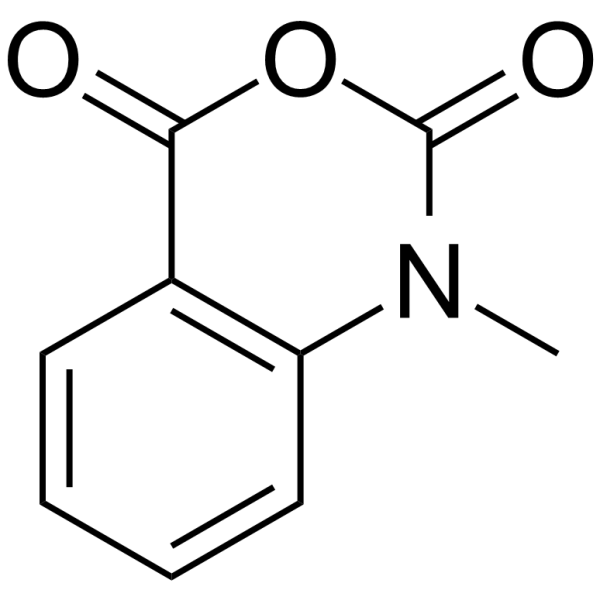
NMIA
|
Others
|
Others
|
|
N-Methylisatoic anhydride (NMIA) is a 2'-OH selective acylation agent of RNAs, and is widely used for
resolving secondary RNA structures using the SHAPE (Selective 2'-Hydroxyl Acylation Analyzed by Primer Extension) technology .
|
-
-
- HY-D0971
-
|
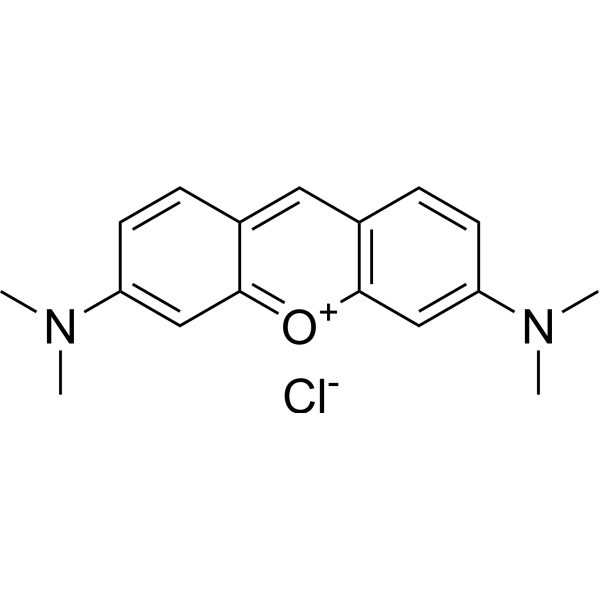
Pyronine G; C.I. 45005
|
DNA Stain
|
Others
|
|
Pyronin Y (Pyronine G) is a cationic dye that intercalates RNA and has been used to target cell structures including RNA, DNA and organelles. Pyronin Y forms fluorescent complexes with double-stranded nucleic acids (especially RNA) enabling semi-quantitative analysis of cellular RNA. Pyronin Y can be used to identify specific RNA subspecies of ribonuclear proteins complexes in live cells .
|
-
-
- HY-W008990
-
|
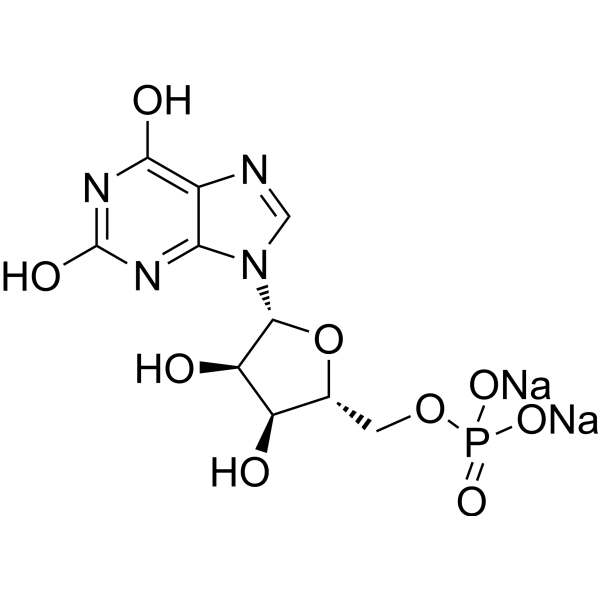
5'-Xanthylic acid sodium salt
|
DNA/RNA Synthesis
|
Metabolic Disease
|
|
Xanthosine 5'-monophosphate sodium salt (5'-Xanthylic acid sodium salt) is an intermediate in purine metabolism. Xanthosine 5'-monophosphate sodium salt can be used for genetic code, nucleic acid structure, and DNA, RNA and protein synthesis research .
|
-
-
- HY-145982
-
|
|
Nucleoside Antimetabolite/Analog
|
Others
|
|
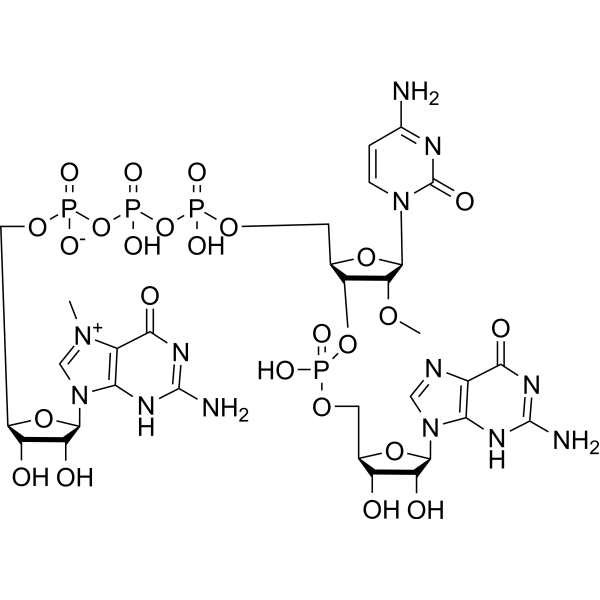
m7GpppCmpG, an oligonucleotide, is an M 7GpppNpG trinucleotide cap analogue. m7GpppCmpG can be used as a chemical tool enabling manufacturing of RNA featuring either cap 0 or cap 1 structures .
|
-
-
- HY-145981
-
|
|
Nucleoside Antimetabolite/Analog
|
Others
|
|
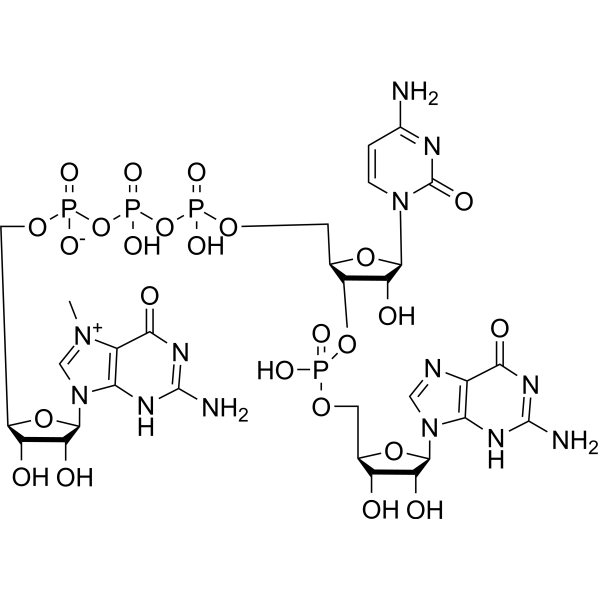
m7GpppCpG, an oligonucleotide, is an M 7GpppNpG trinucleotide cap analogue. m7GpppCpG can be used as a chemical tool enabling manufacturing of RNA featuring either cap 0 or cap 1 structures .
|
-
-
- HY-145979
-
|
|
Nucleoside Antimetabolite/Analog
|
Others
|
|
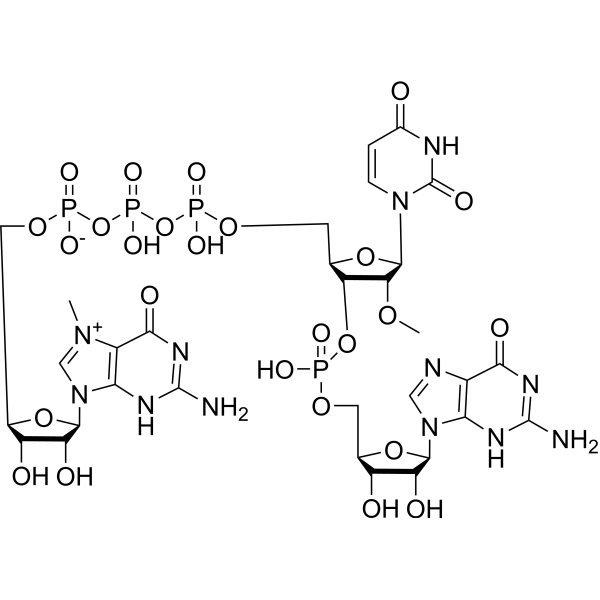
m7GpppUmpG, an oligonucleotide, is an M 7GpppNpG trinucleotide cap analogue. m7GpppUmpG can be used as a chemical tool enabling manufacturing of RNA featuring either cap 0 or cap 1 structures .
|
-
-
- HY-145980
-
|
|
Nucleoside Antimetabolite/Analog
|
Others
|
|
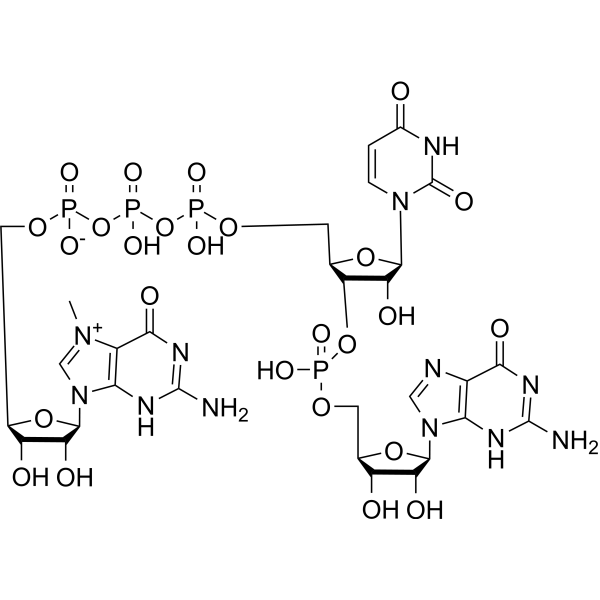
m7GpppUpG, an oligonucleotide, is an M 7GpppNpG trinucleotide cap analogue. m7GpppUpG can be used as a chemical tool enabling manufacturing of RNA featuring either cap 0 or cap 1 structures .
|
-
-
- HY-113061
-
-
-
- HY-145442
-
|
|
Others
|
Others
|
|
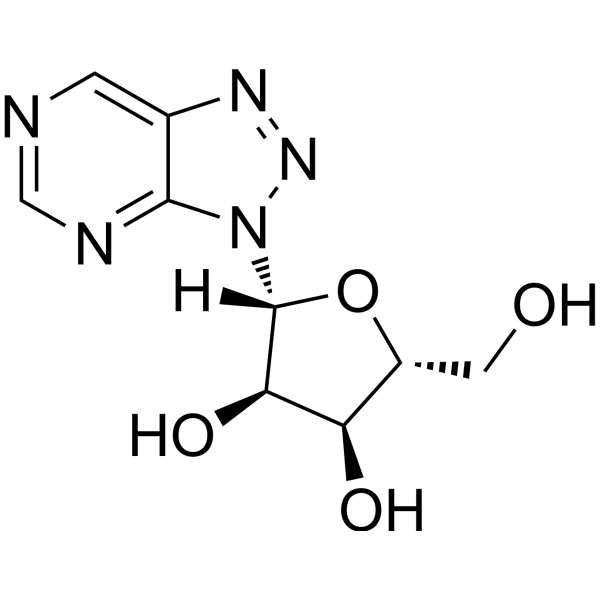
8-Azanebularine, a compound with hydrogen in place of the C6 amino group, inhibits the ADAR2 reaction at high concentrations (IC50=15 mM). 8-Azanebularine is incorporated into an RNA structure recognized by human ADAR2 results in high-affinity binding (KD=2 nM). 8-Azanebularine can be used for the research of ADAR-catalyzed RNA-editing reaction .
|
-
-
- HY-14776
-
|
CX-3543
|
DNA/RNA Synthesis
|
Cancer
|
|
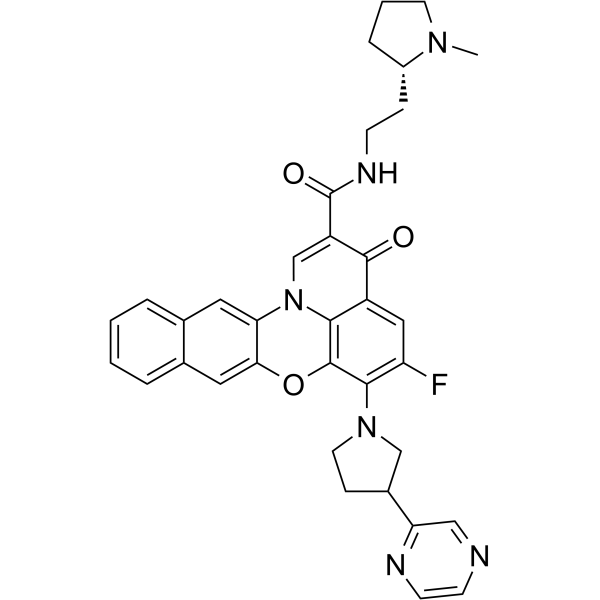
Quarfloxin (CX-3543), a fluoroquinolone derivative with antineoplastic activity, targets and inhibits RNA pol I activity, with IC50 values in the nanomolar range in neuroblastoma cells. Quarfloxin disrupts the interaction between the nucleolin protein and a G-quadruplex DNA structure in the ribosomal DNA (rDNA) template .
|
-
-
- HY-113061S
-
-
-
- HY-120118
-
|
ML246
|
DNA/RNA Synthesis
|
Cancer
|
|
Metarrestin (ML246) is an orally active, first-in-class and specific perinucleolar compartment inhibitor. Metarrestin disrupts the nucleolar structure and inhibits RNA polymerase (Pol) I transcription, at least in part by interacting with the translation elongation factor eEF1A2. Metarrestin blocks metastatic development and extends survival in mouse cancer models .
|
-
-
- HY-W728005
-
|
|
SARS-CoV
|
Infection
|
|
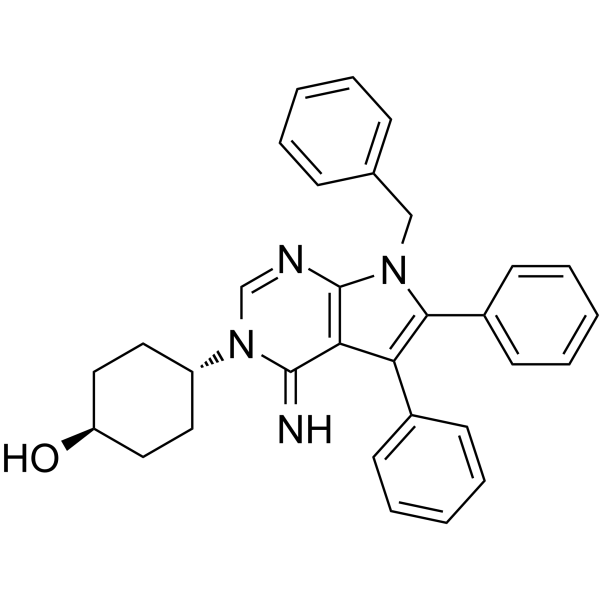
Covidcil-19 (compound C5) avidly binds to the revised attenuator hairpin structure of the SARS-CoV-2 frameshifting element (FSE) with a Kd of 11 nM. Covidcil-19 stabilizes the hairpin’s folded state and impairs frameshifting in cells. Covidcil-19 reduces frameshifting efficiency of the SARS-CoV-2 FSE and does not affect SARS-CoV-2 FSE RNA levels. Covidcil-19 inhibits a process essential for SARS-CoV-2 viral propagation .
|
-
-
- HY-100008
-
|
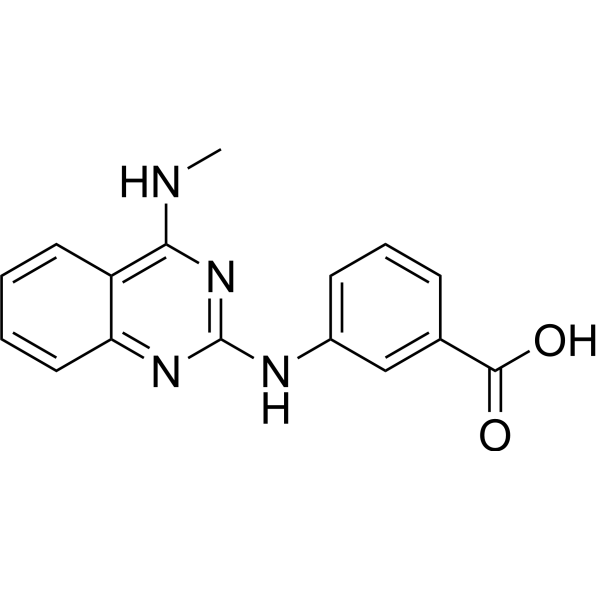
NIK333
|
RAR/RXR
SphK
Autophagy
HCV
|
Infection
Cancer
|
|
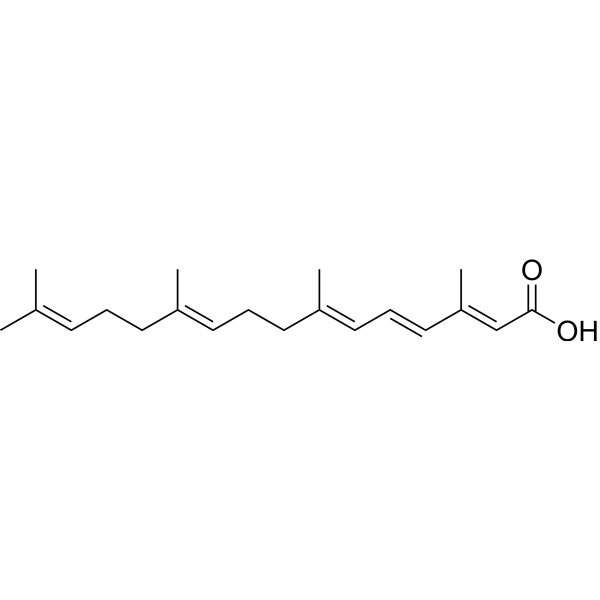
Peretinoin is an oral acyclic retinoid with a vitamin A-like structure that targets retinoid nuclear receptors such as retinoid X receptor (RXR) and retinoic acid receptor (RAR). Peretinoin reduces the mRNA level of sphingosine kinase 1 (SPHK1) in vitro by downregulating a transcription factor, Sp1 . Peretinoin prevents the progression of non-alcoholic steatohepatitis (NASH) and the development of hepatocellular carcinoma (HCC) through activating the autophagy pathway by increased Atg16L1 expression . Peretinoin inhibits HCV RNA amplification and virus release by altering lipid metabolism with a EC50 of 9 μM .
|
-
-
-
HY-L005
-
|
|
1225 compounds
|
|
Epigenetics refers to changes in phenotype that are not rooted in DNA sequence. Many types of epigenetic processes have been identified, including DNA methylation, alteration in the structure of histone proteins and gene regulation by small noncoding microRNAs. Modification of DNA, protein, or RNA, resulting in changes to the function and/or regulation of these molecules, without altering their primary sequences, reveals the complexities of cellular differentiation, embryology, the regulation of gene expression, aging, cancer, and other diseases.
MCE provide a unique collection of 1225 epigenetics-related compounds that can be used in the research of the related diseases.
|
-
-
HY-L073
-
|
|
283 compounds
|
|
Hepatitis C virus (HCV) is a hepatotropic enveloped positive- strand RNA virus (family Flaviviridae) that infects the parenchymal cells of the liver. HCV infection is a significant public health burden. Globally, an estimated 71 million people have chronic hepatitis C virus infection. A significant number of those who are chronically infected will develop cirrhosis or liver cancer. To date, there is no vaccine against HCV, and combination pegylated alpha interferon (pIFN-) and ribavirin, the main standard-of-care treatment for HCV, is effective in only a subset of patients and is associated with a wide spectrum of toxic side effects and complications. More recently, new therapeutic approaches that target essential components of the HCV life cycle have been developed, including direct-acting antiviral (DAA) that specifically block a viral enzyme or functional protein and host-targeted agents (HTA) that block interactions between host proteins and viral components that are essential to the viral life cycle. However, the genetic diversity of HCV viruses and the stage of liver disease (i.e., cirrhosis) are revealing themselves as obstacles for effective, pan-genotypic treatments. There still exists a need for the discovery and development of new HCV inhibitors. In particular, since the future of HCV therapy will likely consist of a cocktail approach using multiple inhibitors that target different steps of infection, new antivirals targeting all steps of the viral infection cycle.
MCE offers a unique collection of 283 compounds with identified and potential anti-HCV activity. MCE Anti- Hepatitis C Virus Compound Library is a useful tool for discovery new anti-HCV drugs and other anti-infection research.
|
-
-
HY-L048
-
|
|
339 compounds
|
|
The high rates of morbidity and mortality caused by fungal infections are associated with the current limited antifungal arsenal and the high toxicity of the compounds. Additionally, identifying novel drug targets is challenging because there are many similarities between fungal and human cells. The most common antifungal targets include fungal RNA synthesis and cell wall and membrane components, though new antifungal targets are being investigated. Nonetheless, fungi have developed resistance mechanisms, such as overexpression of efflux pump proteins, overexpression and changes in drug targets and biofilm formation, emphasizing the importance of discovering new antifungal drugs and therapies. Due to the limited antifungal arsenal, researchers have sought to improve treatment via different approaches, such as the combination of antifungal drugs, development of new formulations for antifungal agents and modifications to the chemical structures of traditional antifungals, etc.
MCE offers a unique collection of 339 compounds with validated antifungal activities. MCE antifungal compound library is an effective tool for drug repurposing screening, combination screening and biological investigation.
|
-
-
HY-L155
-
|
|
470 compounds
|
|
Mitochondria, as the main place of energy supply in life, is essential to maintain normal life activities. Mitochondrial dysfunction is associated with common diseases, such as cardiovascular diseases, neurodegenerative diseases, diabetes and cancer. The heart, brain and liver rely heavily on mitochondrial function as the main organs for drug metabolism. In addition, mitochondria is also a target of many drugs, some of which induce organotoxicity by inducing mitochondrial toxicity.
MCE contains 470 mitochondrial toxic compounds, which can be used as tool compounds for drug development, organ toxicity and disease mechanism research.
|
-
-
HY-L004
-
|
|
2001 compounds
|
|
DNA is prone to numerous forms of damage that can injure cells and impair fitness. Cells have developed an array of mechanisms to repair these injuries. Proliferating cells are especially vulnerable to DNA damage due to the added demands of cellular growth and division. Cell cycle checkpoints represent integral components of DNA repair that coordinate cooperation between the machinery of the cell cycle and several biochemical pathways that respond to damage and restore DNA structure. By delaying progression through the cell cycle, checkpoints provide more time for repair before the critical phases of DNA replication, when the genome is replicated, and of mitosis, when the genome is segregated. Loss or attenuation of checkpoint function may increase spontaneous and induced gene mutations and chromosomal aberrations by reducing the efficiency of DNA repair.
MCE owns a unique collection of 2001 cell cycle/DNA damage-related compounds which can be used in the research of the same.
|
| Cat. No. |
Product Name |
Type |
-
- HY-D0971
-
|
Pyronine G; C.I. 45005
|
Fluorescent Dyes/Probes
|
|
Pyronin Y (Pyronine G) is a cationic dye that intercalates RNA and has been used to target cell structures including RNA, DNA and organelles. Pyronin Y forms fluorescent complexes with double-stranded nucleic acids (especially RNA) enabling semi-quantitative analysis of cellular RNA. Pyronin Y can be used to identify specific RNA subspecies of ribonuclear proteins complexes in live cells .
|
-
- HY-D1408
-
|
DMTr-4'-Methyluridine-CED-TBDMS phosphoramidite
|
Oligonucleotide Labeling
|
|
DMTr-4'-Me-U-CED-TBDMS phosphoramidite (DMTr-4'-Methyluridine-CED-TBDMS phosphoramidite), a dye reagent for oligonucleotide labeling, can be used for the research of applications in RNA therapeutics, RNA aptamers, and ribozymes for elucidating RNA structure. DMTr-4'-Me-U-CED-TBDMS phosphoramidite represents a probe with wide utility for elucidation of RNA structure .
|
-
- HY-D1409
-
|
DMTr-4'-F-uridine-CED-TBDMS phosphoramidite
|
Oligonucleotide Labeling
|
|
DMTr-4'-F-U-CED-TBDMS phosphoramidite (DMTr-4'-F-uridine-CED-TBDMS phosphoramidite), a dye reagent for oligonucleotide labeling, can be used for the research of applications in RNA therapeutics, RNA aptamers, and ribozymes for elucidating RNA structure. DMTr-4'-F-U-CED-TBDMS phosphoramidite represents a probe with wide utility for elucidation of RNA structure .
|
| Cat. No. |
Product Name |
Type |
-
- HY-153108
-
|
ARCA cap
|
Biochemical Assay Reagents
|
|
3'-O-Me-m7G(5')ppp(5')G (ARCA cap), anti-reverse cap analog, has a special RNA cap structure. Is a common feature of the mRNA of some RNA viruses and eukaryotes. RNA cap structures serve as signals for translation initiation .
|
| Cat. No. |
Product Name |
Category |
Target |
Chemical Structure |
| Cat. No. |
Product Name |
Chemical Structure |
-
- HY-113061S
-
|
|
|
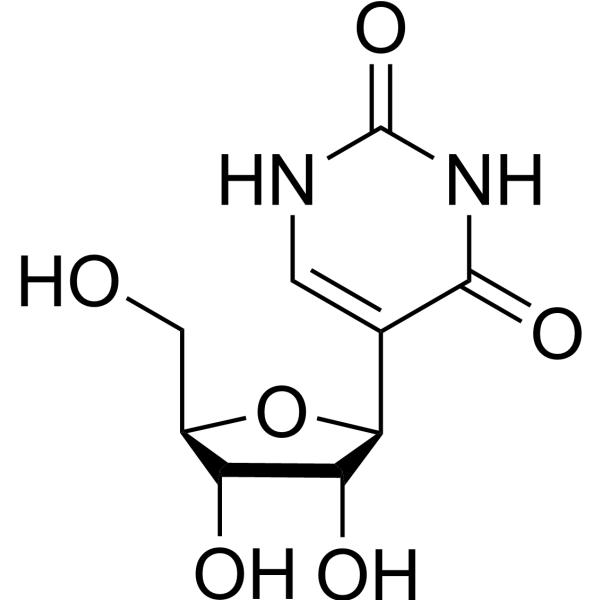
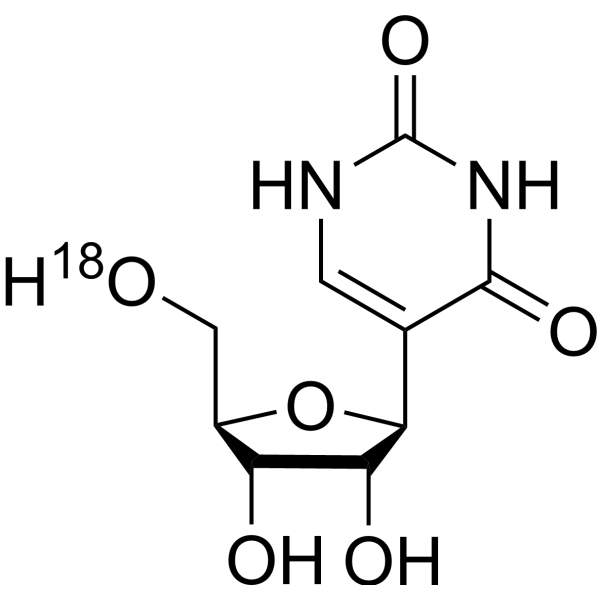
Pseudouridine- 18O is the 18O labeled Pseudouridine (HY-113061). Pseudouridine is an isomer of the nucleoside uridine, and the most abundant modified nucleoside in non-coding RNAs. Pseudouridine in rRNA and tRNA can fine-tune and stabilize the regional structure and help maintain their functions in mRNA decoding, ribosome assembly, processing and translation.
|
-

| Cat. No. |
Product Name |
|
Classification |
-
- HY-103006
-
NAI-N3
5 Publications Verification
|
|
Azide
|
|
NAI-N3 is a RNA acylation reagent that enables RNA purification. NAI-N3 is a dual-function SHAPE (selective 2-hydroxyl acylation and profiling experiment) probe (RNA structure probe and enrichment) .
|
-
- HY-163213
-
|
|
|
Azide
|
|
Psoralen-triepthylene glycol azide is a compound used to probe the structure and conformation of RNA in living cells, using matching RNA crosslinking and deep sequencing or comradery methods .
|
Your information is safe with us. * Required Fields.
Inquiry Information
- Product Name:
- Cat. No.:
- Quantity:
- MCE Japan Authorized Agent:



























